Synthedia creates synthetic LC-MS/MS data that mimics real data but with a composition that is exactly known. Currently, synthedia support the creation of Data-Independent Acquisition (DIA) style data wherein fixed, large m/z windows are sequentially isolated for fragmentation. We have focused on creating DIA data to date since the complexity of the analysis preformed by processing tools is substantial and the impact of different acquisition methodologies on the eventual outcome is somewhat more difficult to predict.
Required input files
To simulate DIA data with Synthedia, a set of peptide precursors and fragments must be supplied. This can be given as either a Prosit library, or MaxQuant search results files from the analysis of data-dependent acquisition (DDA) data.
Prosit libraries
Prosit is a machine learning-based application that predicts MS/MS fragmentation patterns for input peptide sequences. Input sequences can be arbitrary and do not necessarily need to originate from any specific organism or protein. Prosit will generate an output file that contains predicted abundances for peptide sequence ions.
To create a Synthedia-compatible Prosit spectral library:
- Go to https://www.proteomicsdb.org/prosit/
- Navigate to 'Spectral Library'
- Upload your target peptide list as described in the Prosit documentation
- IMPORTANT: under the 'Output format' header (just prior to submitting the Prosit task), ensure that Generic Text is selected
Once processing is complete, the resulting file can be used with Synthedia
When Prosit inputs are given, Synthedia models peptide precursor abundances using a Gaussian distribution of Log2 instnsities which approximates the disributions typically observed upon anlysis of real data. The parameters of this distribution (mean and standard deviation) can be specified with the --prosit_peptide_abundance_mean and --prosit_peptide_abundance_stdev parameters which have defaults of 22 and 3 respectively.
MaxQuant 'txt' directories
As an alternative to Prosit, Synthedia can read and simulate DIA data based upon the MaxQuant processing results of a file acquired using a Data-Dependent Acquisition (DDA) strategy. In this case, peptide fragment ions are generated based on the matched ions for a PSM reported in the MaxQuant msms.txt file from the Masses and Intensities columns. Note: these are only those fragment ions that MaxQuant assigns as matching a given peptide - they may not necessarily provide a 'full' sequence coverage and may not be correctly assigned in some cases.
Synthedia offers options to filter reverse and contaminant peptides as well as filter PSMs by Posterior Error Probability (PEP) values.
Optional input files
Decoy signals file
LC-MS/MS analysis of bottom-up proteomics samples can be complicated by the presence of ions derived from non-peptide sample contaminants. To mimic this, decoy ions can be simulated together with peptide signals by specifying a decoy database in NIST '.msp' format. See the section 'NIST Text Format of Individual Spectra' in this document for details on the .msp format.
Custom .msp files can be specified or pre-prepared files can be dowloaded from MS-DIAL. In this online implementation of Synthedia, if decoys are requested but no MSP file is given, the "All public MS/MS (13,303 unique compounds)" file from MS-DIAL is used.
The number of decoy peaks to simulate, as well as the maximum number of fragments to simulate per decoy, can be specified throught the command line arguments num_decoys and simulate_top_n_decoy_fragments.
DIA acquisition schema file
DIA-LC-MS/MS data can be acquired in many ways. The default invocation of synthedia creates non-overlapping, 30 Th windows between m/z 350 and m/z 1600. To simulate data using different DIA acquisition strategies, a file defining the acquisition schema can be supplied. An example acquisition schema file and blank templates can be downloaded from the resources tab.
Compatibility of Synthedia mzML files with other software
The mzML files generated with Synthedia have been tested with:
- DIA-NN
- EncyclopeDIA
- DIAUmppire
- Skyline
- MSConvert
- OpenMS/ToppView
The mzML files are known to not be compatible with MaxQuant (at least as at MaxQuant Version 2.0.3.0)
Notes about the resulting data
Peptide vs protein abundances
In a real experiment, all peptides from an up-regulated protein should be observed with increased abundance compared to a control group. Synthedia models ion abundances at the peptide level only. This means that, in the case of a two-group simulation, peptides from the same protein may have very different (even opposing) directions of abundance change between groups. This is primarily because the Prosit input type (which is preferred) does not contain mappings between peptides and input proteins and we wished to maintain the ability to simulated arbitrary peptide data.
As a result of this, if mzML files generated with synthedia are analysed with DIA analysis software, the main comparison in abundances should be made at the peptide level.
Synthetic vs real data
While we have endeavoured to provide a range of options that allow for simulation of a broad range of chromatographic and mass spectral variables, many experimental processes are not modelled which will cause deviations between simulated and real data. As such, users are warned that simulations with Synthedia do not completely re-create experimental complexity and should be used as an investigational tool only.
As an example, Synthedia offers the capability to simulate the same set of precursors as if acquired on different length chromatographic gradients. This means that data acquired on a lengthy gradients could be reconstructed to approximate acquisition on shorter gradients. In these cases, Synthedia simply models spectra containing signals from many peptides as a simple superposition of their individual signals. In reality however, ion supression effects would result in data that may look substantially different from that which would be produced if a comparable experimental workflow were to be executed.
Post-translational modifications
Synthedia currently has very limited support for simulating peptides with post-translational modifications. Currently, all cysteine residues are assumed to be modified by carbamidomethylation. No other post-translational modifications are supported.
Simulation time requirements
Depending on your dataset and parameters, simulations can be quite time consuming. For this reason, some restrictions are placed on the data that can be simulated with this web implementation of Synthedia. These are:
- The maximum number of groups that can be simulated is 3
- The maximum number of replicates per group that can be simulated is 6
If you wish to run a simulation that exceeds these, either download and run Synthedia locally, or contact us to arange a collaboration.
As a rough guide, the simulation of 37,000 precursors in full centroid mode takes approximately 15 minutes per mzML file.
Reporting Issues
If you notice bugs or have suggestions for improvements, please contact us here or email us
Synthedia output file sets
A symthedia data simulation produces a range of output files. These are described below:
| File |
Description |
| mzML file(s) |
One or more mzML file containing the simulated DIA-LC-MS/MS data. The file name suffix _group_X_sample_Y indicate the group to which the file belongs in the case that multiple treatment groups are simulated. |
| peptide_table.tsv |
Tab-separated value file that defines the precursors simulated and the properties (retention time, m/z, abundances) in the simulated mzML files. See below for a detailed description of the contents of this file. |
| peptides.pickle |
File containing simulated peptide metadata. This file can be used to repeat an earlier simulation. Since some paramaters are randomly generated (Eg. peptide abundance differences between treatment groups), repeating a simulation from this file will be slightly different to re-running a simulation with the same parameters from scratch. |
| simulation_args.yaml |
Human readable file containing input arguments used to start the simulation. |
| assembly.log |
Text file containing logging data about the simulation. |
Description of peptide_table.tsv
The columns in the peptide_table.tsv output file are described below:
| Column |
Description |
| Protein |
The protein (or protrin group) to which a precursor belongs. Note that this will be 'none' for Prosit input types. |
| Sequence |
The peptide sequence of the simulated ion. Note that all Cys residues are assumed to be carbamidomethylated. |
| Intensity |
The precursor intensity specified in the input data set. in the case of Prosit inputs, this is arbitrarily set to 100,000,000. In the case of MaxQuant inputs, this is taken as the value in the 'Intensity' column of the 'evidence.txt' file. |
| m/z |
The precursor m/z value taken from the input data files. In the case of Prosit inputs, this is taken as the value in the PrecursorMz column. In the case of MaxQuant inputs, this is taken as the value in the 'm/z' column of the 'evidence.txt' file. |
| Charge |
The precursor charge. In the case of Prosit inputs, this is taken as the value in the PrecursorCharge column. In the case of MaxQuant inputs, this is taken as the value in the 'Charge' column of the 'evidence.txt' file. |
| Mass |
The neutral mass of the peptide. In the case of Prosit inputs, this is taken as the PrecursorMz column value multiplied by the PrecursorCharge column value minus the PrecursorCharge multiplied by 1.007276. In the case of MaxQuant inputs, this is taken as the value in the 'Mass' Column. |
| Experimental RT |
The retention time of the precursor in the input data file. In the case of MaxQuant inputs, this is taken as the value in the 'Retention time' column multiplied by 60. In the case of Prosit inputs, this is taken as the value in the 'iRT' column plus the minimum value in the 'iRT' column |
| Synthetic RT group_X_sample_Y |
The retention time (in seconds) of the precursor in the simulated output data. |
| Synthetic RT Start group_X_sample_Y |
The retention time of the first mass spectrum (MS1 or MS2) in which a precursor is observed in the synthetic data of sample Y of group X. |
| Synthetic RT End group_X_sample_Y |
The retention time of the last mass spectrum (MS1 or MS2) in which a precursor is observed in the synthetic data in sample Y of group X. |
| Synthetic theoretical m/z 0 |
The theoretical m/z value of the monoisotopic peak of a precursor in the synthetic data. |
| Synthetic m/z group_X_sample_Y |
The simulated m/z value of the monoisotopic peak of a precursor in the synthetic data. This will be different from the theoretical m/z value if a ppm error is included in the simulation. |
| Synthetic m/z ppm error group_X_sample_Y |
The part-per-million error between the theoretical peptide m/z value and the value simulated in the synthetic data. |
| Total abundance group_X_sample_Y |
Sum of the intensities of all MS1 data points attributable to a given precursor in sample Y of group X. |
| Peak height group_X_sample_Y |
Maximum intensity of all MS1 data points attributable to a given precursor in sample Y of group X. |
| Offset group_X_sample_Y |
Log2 value of the abundance offset between the value in the 'Intensity' column and the simulated intensity for sample Y of group X. |
| Found in group_X |
Indicates whether the precursor is present in group X. A value of '1' indicates that the precursor is present in one or more samples of group X. A value of '0' indicates that the precursor is not present in any sample of group X. |
| Found in group_X_sample_Y |
Indicates whether the precursor is present in sample Y group X. A value of '1' indicates that the precursor is present in group X sample Y. A value of '0' indicates that the precursor is not present in group X sample Y. |
| MS1 chromatographic points group_0_sample_0 |
Number of MS1 chromatographic points in which a precursor is observed in the simulated data. |
| MS2 chromatographic points group_0_sample_0 |
Number of MS2 chromatographic points in which a precursor is observed in the simulated data. |
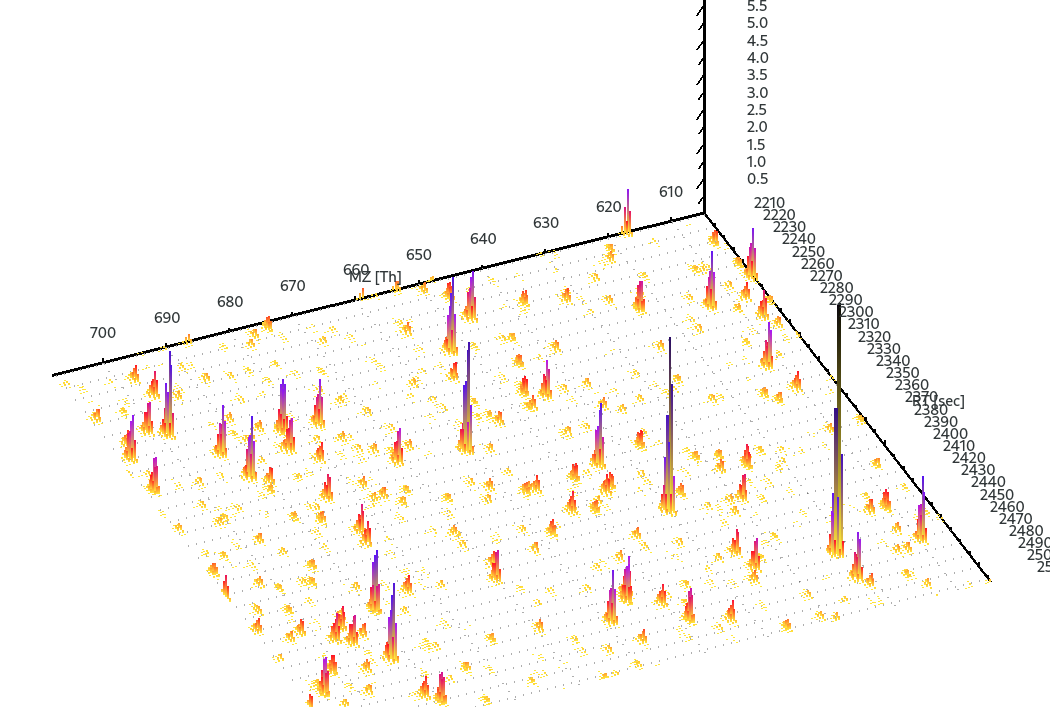
Viewing mzML files
The mzML files produced with Synthedia can be viewed in many different freely available software such as TOPPView package which is part of OpenMS.